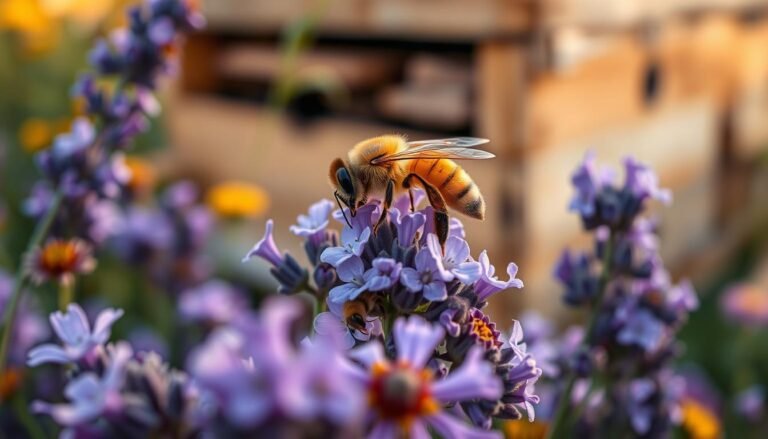
This scientific review shows how heritable regulation of gene expression links a shared genome to distinct queen and worker outcomes. We outline key mechanisms—DNA methylation, histone modification, and non-coding RNAs—that shape larval fate and adult behavior.
Bees reveal a unique methylome: low global 5mC but exon-enriched methylation that stabilizes transcription and affects alternative splicing. This pattern helps produce caste- and tissue-specific expression programs without changing DNA sequence.
Dietary cues matter. Royal jelly supplies 10-HDA and royalactin-like proteins that act on chromatin and signaling to bias larvae toward a queen path. Adult worker roles also show plasticity, with reversible methylation shifts tied to nurse-forager transitions and pheromone signals.
Methodological advances—methylome sequencing, ChIP-seq, RNAi, and pharmacological tools—now map these processes precisely. We close by noting applied implications for U.S. beekeeping, from queen longevity to chemical exposures affecting colony health.
Key Takeaways
- Shared genes can yield queen and worker outcomes through heritable regulation of gene expression.
- DNA methylation in bees is exon-enriched and influences transcription stability and splicing.
- Royal jelly contains bioactives that alter chromatin and bias larval fate toward queens.
- Adult social roles are flexible and show reversible methylation tied to behavior and pheromones.
- New genomic and epigenomic tools enable precise mapping of mechanisms in Apis mellifera.
- Findings have practical relevance for queen management, nutrition, and chemical risk in U.S. apiaries.
Abstract and Scope of This Scientific Review Article
This review synthesizes current data and methods that reveal how chromatin, chemical marks, and short RNAs coordinate gene networks to produce distinct queen and worker phenotypes from the same genome.
We summarize aims across molecular to colony scales. Topics include dna methylation, histone marks, and non-coding RNAs as core mechanisms for regulation of genes and gene expression.
The article highlights nutritional inputs such as royal jelly as upstream cues that remodel chromatin and bias developmental trajectories. Case studies cover nurse-forager transitions, pheromonal signaling, organ-level outcomes, and immunity.
Methodological strengths and limits are discussed, with attention to methylome sequencing and ChIP-seq. For a focused view on sequencing approaches see methylome and ChIP-seq datasets.
Applied relevance is emphasized for U.S. beekeeping: insights into queen quality, nutrition strategies, and exposure management link basic mechanisms to practical outcomes.
“This synthesis positions chromatin-based regulation as a unifying framework for phenotype plasticity across social insects.”
Why Epigenetics Matters in Honeybees: From Genome to Phenotype
Molecular marks translate environmental signals into stable patterns of gene activity. These marks act without changing the DNA sequence, yet they can be passed through cell divisions and, sometimes, between generations.
Defining regulatory control and plastic outcomes
Epigenetics describes reversible, heritable regulation that tells a gene when and where to act. This control lets a single genome produce multiple adult forms via phenotypic plasticity.
Reversible marks versus sequence change
Unlike mutations, chemical tags such as methylation, histone modifications, and small RNAs can flip on short notice. That rapid flip supports flexible responses to diet, season, or social cues in social insects.
- Mechanisms: DNA methylation affects exon usage and stable expression.
- Integration: Histone marks and ncRNAs relay external signals into gene expression outcomes.
- Adaptive value: Reversible marks enable behavioral shifts and can alter phenotypic variance across a species.
Honeybee Caste Biology: Queens, Workers, and Developmental Divergence
A single genome gives rise to two distinct adult forms with unique roles and lifespans.
Queens are large, highly fertile, and long-lived. They invest in egg production and reproductive success.
Workers are smaller, typically sterile, and specialized for foraging, brood care, and colony defense. Their morphology supports these tasks.
Shared genome, contrasting traits
Both castes start from genetically identical eggs. Diet—royal versus worker jelly—sets each larva on a different path.
Early larval windows are critical: timing of feeding and nutritional signals lock in tissue-specific gene programs.
| Trait | Queen | Worker |
|---|---|---|
| Body size | Large | Smaller |
| Reproductive organs | Well-developed ovaries | Reduced ovaries |
| Glandular development | Reproductive glands | Hypopharyngeal and mandibular glands |
| Lifespan | Years | Weeks to months |
These differences reflect distinct gene expression profiles shaped by chromatin and chemical marks during larval growth.
Worker behavior shows plasticity: shifts in colony demography can reprogram tasks and roles, revealing flexible adult gene regulation.
Fitness trade-offs are clear: queens prioritize reproduction, while workers optimize labor and colony defense. Together, these roles make the colony a superorganism that maximizes collective fitness.
Epigenetics in honeybee caste development
Larval feeding patterns set a molecular program that steers future anatomy and behavior. Small shifts in timing or content of food reach chromatin and alter how the genome is used.

Nutritional cues as upstream signals for epigenetic programming
Diet quality and timing act as primary environmental signals. Sustained royal jelly feeding supplies bioactive factors that change dna methylation and histone acetylation in larvae destined for reproduction.
Worker-fed larvae receive different glandular secretions. That contrast sends divergent instructions to gene networks and sets alternative growth paths.
Environment–genome interaction shaping caste fate
Early larvae form an inherited bridging phenotype — a transient, nutrition-sensitive state primed to read external cues. This state channels gene-by-environment interactions toward conditional outcomes.
Epigenetic mechanisms capture environmental information and modulate gene expression. The genome provides boundary conditions; epigenetic “software” integrates context and executes control programs.
“Dietary signals are translated into lasting molecular marks that align individuals with colony needs.”
- Worker-controlled feeding ensures reliable signal transmission across generations.
- Responses are adaptive: larvae align traits with predicted colony demands.
- Many marks remain reversible, allowing plastic shifts if social or environmental conditions change.
Next: we examine core methylation and chromatin mechanisms that link diet to gene regulation at the exon and tissue levels.
DNA Methylation: Core Mechanism Linking Diet to Development
Gene-body methylation concentrates across exons and guides how transcripts are spliced and used. This exon-centered pattern is a hallmark of the species and helps create isoform diversity while stabilizing steady-state expression.
Gene-body marks and transcript processing
Exonic methylation correlates with alternative splicing. Genes with high exon methylation often show stable output and controlled isoform ratios. Low-CpG promoters behave differently, with methylation more likely to repress transcription.
Maintenance versus de novo enzymes
DNMT1 preserves methylation during cell division. DNMT3 writes new marks during critical larval windows. Diet can modulate both enzymes, linking nutrition to methylation setting.
Tissue, stage, and germline dynamics
Levels vary by tissue: fat body often shows higher methylation than brain. Larval and pupal stages follow distinct trajectories; queen-directed larvae show lower methylation early, then diverge later from worker patterns.
- Oocytes carry higher methylation than sperm, affecting early embryonic genomes.
- Methylation acts as a transcriptional stabilizer while permitting plastic change via splicing control.
“Methylation maps provide a bridge from diet to organ-level and behavioral gene networks.”
Royal Jelly, Major Royal Jelly Proteins, and Epigenetic Targets
A potent mix of proteins and small molecules in royal jelly rewires signaling and chromatin during early larval windows.
Composition and gene origins. Royal jelly is rich in Major Royal Jelly Proteins (MRJPs), produced from a nine-gene array derived from the yellow gene family. These proteins likely helped the evolution of social traits by changing nutrition and behavior.
MRJPs and the yellow gene family expansion
MRJPs act as both nutrients and signaling ligands. Their gene expansion provides multiple paralogs with distinct expression and functional roles during larval growth.
10-HDA, DNMT3A and histone targets
10-HDA is a fatty acid that inhibits histone deacetylase activity, notably affecting HDAC3 and keeping histone marks acetylated. This open-chromatin state supports active transcription and reduced de novo methylation via DNMT3A inhibition, biasing methylation levels toward queen pathways.
Royalactin and growth signaling
Royalactin triggers MAPK and p70 S6K and elevates juvenile hormone. These signals accelerate body growth and ovary maturation; effects appear conserved across species, including Drosophila.
- Worker versus queen diets differ: worker jelly contains more plant-derived phenolic acids found in honey-pollen mixes, with implications for immunity and detoxification.
- Mandibular and hypopharyngeal gland secretions determine precise nutrient and epigenetic cue profiles delivered to larvae.
Timing and dose matter. Sustained royal jelly feeding during critical windows consolidates queen-like expression programs. Dosage and timing shape organogenesis and reproductive capacity, giving workers control over caste trajectories.
Dietary Determination of Caste: Timing, Content, and Control
Colony nurses act as nutritional gatekeepers, tailoring meals that bias gene activity and future roles. Worker behavior sets both the volume and biochemical content delivered to each larva. This precise control determines whether a larva follows a queen or worker path.
Worker gland roles
Hypopharyngeal and mandibular glands produce distinct jellies. Royal-like secretions contain proteins and 10-HDA that keep chromatin open and favor queen traits. Worker jelly has different sugars and plant-derived compounds that bias toward labor roles.
Critical early window
Early larvae (about 48 hours) form a responsive bridging phenotype. Feeding during this narrow period sets dna methylation and histone acetylation states that change genes and expression levels long term.
Colony-level control matters. Food stores and nurse demographics limit how many extended royal diets the colony can sustain. Reduced or altered feeding pushes larvae toward worker outcomes; continuous royal provisioning is required to promote queens rather than cause nutritional castration.
“Nutrition delivered by workers is the proximate switch that links food to lasting gene-regulatory programs.”
Next, we trace how those diet-driven expression patterns shape organ growth and reproductive capacity.
From Differential Gene Expression to Organ-Level Outcomes
Early nutritional cues reshape gene networks that prioritize reproductive tissues over somatic ones.
Diet-driven shifts produce clear transcriptomic signatures that bias ovary growth in queen-destined larvae. Royal jelly–activated insulin signaling raises juvenile hormone and rapidly increases cell proliferation and resource allocation to the ovary. These hormonal changes suppress formation of worker-specific glands and pollen-carrying structures.
At the molecular level, methylation and histone acetylation coordinate metabolic subnetworks that support fast body growth. Epigenetic marks alter expression levels for genes involved in nutrient transport, lipid storage in the fat body, and ribosomal proteins for growth.
Trade-offs are central: larvae on worker diets receive plant phenolics that upregulate detox and immune genes. Queens sacrifice some immunity for reproductive investment, producing a reproductive monopoly that supports colony fitness and division of labor.
Alternative splicing and isoform choice further tune tissue function. Differential nutrient sensing and signaling thresholds set organogenesis outcomes, while sustained regulation of gene expression links those early changes to adult behavior and plasticity.
“Nutrition and hormone crosstalk translate diet into organ-specific gene programs that shape colony roles.”
Behavioral Plasticity in Adult Workers: Nurse-Forager Transitions
Colonies tune workforce behavior by changing neural gene activity in individual workers. Reversion experiments show that task identity maps to reversible methylation patterns in the brain rather than to age alone. These marks shift when nurses become foragers and can flip back when social context demands.
Key genes with task-linked changes include neurexin I, CREB, and Dnmt3. Changes in expression and methylation of these genes track learning and neural plasticity. Odorant‑binding proteins and sensory receptors also show methylation near splice sites that alters isoform use.
Neural marks, DNMT dynamics, and pheromones
DNMT levels rise or fall with associative learning and task shifts, indicating active dna regulation during role change. Methylation entropy increases under threat, suggesting fast neural reprogramming that supports rapid behavioral shifts.
- QMP suppresses ovary growth and recruits nurse behavior.
- Brood pheromone delays nursing-to-foraging and biases pollen collection.
- Ethyl oleate acts as feedback from foragers to slow premature foraging.
| Signal | Physiological effect | Neural/gene targets |
|---|---|---|
| QMP | Suppresses reproduction, promotes nursing | Dnmt3, neuroendocrine genes |
| Brood pheromone | Delays foraging, biases task allocation | Odorant-binding proteins, CREB |
| Ethyl oleate | Feedback inhibition of early foraging | Learning genes, neurexin I |
“Colony-level chemical cues and neural methylation work together to sustain flexible, fitness‑enhancing labor allocation.”
Non-Coding RNAs in Honeybee Epigenetic Regulation
Long noncoding RNAs (lncRNAs) and small RNAs form a regulatory layer that guides chromatin modifiers to target genes. These transcripts act as scaffolds and guides, shaping gene expression and cell fate during reproductive and neural programs.
Lncov1, Lncov2, and ovarian pathways
Lncov1 and Lncov2 link to ovarian growth and reproductive differentiation. They recruit modifiers to loci that control proliferation and hormone signaling. These lncRNAs help shape which genes drive ovary size and function.
Nb-1: embryonic and adult roles
Nb-1 delays zygotic genome activation in males and later appears in octopaminergic neurons and mushroom body stem cells of the worker brain. This dual role ties early sex-specific dna timing to adult neural circuits that affect behavior.
Small RNAs and sensory splicing
miRNAs and siRNAs fine-tune transcript levels for sensory receptors and synaptic proteins. Head lncRNA shifts, often via Wnt signaling, promote alternative splicing of olfactory receptors during nurse-to-forager shifts.
- Cross-talk: ncRNAs interact with methylation and histone marks to set stable but flexible expression states.
- Cell specificity: Many ncRNAs show cell-type patterns in neural lineages tied to learning and memory.
Emerging datasets now enable functional annotation of bee ncRNAs; see work on functional annotation of bee ncRNAs for deeper resources: functional annotation of bee ncRNAs.
“Non-coding RNAs act as complementary layers to methylation and histone marks, refining behavior and reproductive outcomes.”
Histone Modifications and Chromatin State Dynamics
Histone marks act as rapid translators of nutritional signals, shifting chromatin accessibility and directing which genes are read during early growth windows.
Acetylation of histone tails marks open chromatin and supports active transcription. Removal of those acetyl groups compacts chromatin and blocks access by transcriptional machinery. These shifts occur quickly and vary by tissue and time.
HDAC inhibition by royal jelly and consequences for transcription
Royal jelly contains 10‑HDA, a small fatty acid that inhibits histone deacetylase enzymes, notably HDAC3. By blocking HDAC activity, 10‑HDA sustains acetylation levels and keeps target loci accessible for transcription.
This sustained inhibition pairs with lower DNMT3A activity to reduce new methylation at key loci. The result is coordinated changes in methylation and acetylation that favor queen‑like gene networks over worker‑biased programs.
Transcriptional outcomes include greater accessibility for reproductive and growth-related genes, altered expression of metabolic proteins, and shifts in hormone signaling pathways. Sensitivity to dose and timing is high: brief exposure yields temporary changes, while prolonged exposure helps lock in trajectories at the cell level.
“Histone marks act as rapid responders to nutrition, complementing more stable methylation to implement lasting program shifts.”
| Feature | Effect | Implication |
|---|---|---|
| Histone acetylation | Open chromatin, active expression | Permits activation of growth and reproductive genes |
| HDAC3 inhibition (10‑HDA) | Sustains acetylation, reduces deacetylation | Prolongs accessibility during critical windows |
| Interaction with methylation | Reduced de novo methylation at active loci | Stabilizes gene networks favored by nutrition |
Evolutionary Perspectives: From Darwin’s Special Problem to Kin Selection
The puzzle of sterile helpers pushed evolutionary thinkers to expand selection models beyond single organisms. Darwin noted this challenge, calling it a “special difficulty” for his theory.
Group and kin selection frameworks for eusocial traits
Hamilton’s inclusive fitness formalized how cooperation can evolve when relatedness (r) makes rb > c. This math explains how genes that favor helping can spread across a group of close kin.
Group selection and kin selection offer complementary lenses: selection may act on groups when group-level benefits affect survival and reproduction across the species.
Epigenetic inheritance systems and eusocial “superorganisms”
Heritable methylation and other non-sequence mechanisms let trait states shift rapidly without DNA change. Such metastable epialleles can be shaped by colony-level selection to improve task efficiency.
- Colonies function as a superorganism: queens act as reproductive organs; workers serve somatic roles.
- Environmental stress can expose epigenetic variants that selection then favors at the group level.
- Over time, genes may fix beneficial traits, with methylation sometimes influencing mutation rates and long-term stabilization.
“Phenotypic divergence from identical genomes highlights multi-level selection driving social evolution.”
Heritability and Stability of DNA Methylation in Honeybees
Patrilines frequently carry distinct dna methylation landscapes. Samples from the same father share more methylated sites and show fewer differentially methylated regions (DMRs) than random pairs. This pattern points to stable, heritable marks layered on top of genetic variation.
Patriline effects, conservation, and plasticity
Quantitative surveys report roughly 81% of methylation sites conserved between parents and offspring, with little evidence for wholesale reprogramming across the genome. These conserved sites concentrate in exons and regulatory regions, suggesting functional persistence across generations.
At the same time, developmental stress—such as intensive commercial queen breeding or poor rearing conditions—can increase methylation variance. Some of that variation transmits to offspring, showing that stability coexists with context-dependent changes.
- Layered control: methylation often co-varies with SNPs in male germ cells, indicating integrated genetic-epigenetic architectures that affect gene regulation.
- Tissue specificity: links between methylation and expression can be weak or context dependent, especially across brain and ovary samples.
| Feature | Evidence | Implication |
|---|---|---|
| Intergenerational conservation | ~81% shared methylated sites between parents and offspring | Stable marks can predispose lineage-level traits |
| Patriline signature | Fewer DMRs within patrilines; shared methylation profiles | Heritable landscapes overlay genetic variation |
| Stress-driven plasticity | Breeding and rearing stress increase methylation variance | Adaptive remodeling possible under ecological or artificial pressure |
| SNP–methylation covariation | Male germ cell data show co-variation with SNPs | Genetic context modulates methylation effects on genes |
“Stability of methylation does not exclude adaptive change; both patterns shape developmental robustness and evolutionary potential.”
Methodological note: detecting subtle links between methylation and gene expression requires sensitive designs and matched tissue sampling. For applied beekeeping, conserved methylation offers both predictive value and a caution: artificial practices can alter marks that offspring inherit.
Controversies and Open Questions in Honeybee Epigenomics
Conflicting reports now challenge simple links between methylation and gene expression across tissues. Some studies report thousands of differentially methylated regions (DMRs) without matching changes in transcript levels. This mismatch creates uncertainty about causal roles for marks found in sequencing data.
When methylation correlates—or doesn’t—with expression
Work on the brain and ovary often shows weak or absent correlations between methylation and steady-state expression of nearby genes. Behavioral studies add nuance: nurse‑forager reversions reveal reversible methylation that may not change average transcript levels.
- Possible reasons: gene-body marks may affect splicing rather than total transcript counts.
- Chromatin context and promoter versus exon location shape outcomes.
- Queen presence alters DNMT expression, which complicates causal inference from simple datasets.
What’s needed: time-resolved, cell-type-specific assays and multi-omics (methylome, transcriptome, acetylome) to capture transient regulatory mechanisms. Functional validation of DMRs, dose–response tests for dietary modifiers, and standardized designs will help resolve current contradictions.
“Transient dna marks can matter even when steady-state expression shows little change.”
Methods and Data: Omics Approaches Powering Current Insights
Modern omics now let researchers map chemical marks across entire genomes at single‑base resolution. Whole‑genome bisulfite sequencing profiles 5mC and reveals where methylation concentrates across exons and promoters.
Methylome sequencing, ChIP‑seq, and behavioral reversion designs
ChIP‑seq charts histone marks and transcription factor occupancy to link chromatin state to active genes. Paired methylome and RNA profiling connects marks with transcript-level outputs and functional gene networks.
Behavioral reversion paradigms (nurse↔forager) induce reversible DMRs and show Dnmt3 upregulation after associative learning. Designs that control chronological age separate behavior-linked methylation from age effects.
Assays for DNMT activity and pharmacological perturbations (decitabine, zebularine) test causal roles for DNA methylation. Careful sample stratification—brain, fat body, ovary, and developmental stages—improves inference.
- Statistics: robust DMR calling, methylation entropy, and multi‑factor models are essential.
- Challenges: low global methylation, cell heterogeneity, and the need for single‑cell or nucleus methods.
“Open data and standardized pipelines will enable cross‑lab validation and larger meta‑analyses.”
Applied Implications: Beekeeping Practices and Bee Health in the United States
Beekeepers can use molecular insights to make simple, impactful changes to nutrition and breeding. This helps sustain queen longevity and worker performance while lowering chemical risk to the colony.
Queen longevity, chemical exposures, and sustainable nutrition strategies
Prioritize sustained, high-quality feeding. Ensure colonies have diverse forage and supplemental feeds that mimic natural content to support immunity and balanced gene regulation.
Recognize queen vulnerability. Royal diets lack many plant phenolics found in worker food. Queens can be more sensitive to pesticides and toxins. Adjust treatment timing and minimize chronic exposures.
- Limit stressful commercial rearing practices that raise methylation variance and reduce offspring robustness.
- Promote forage diversity to supply nutrients that support detoxification and immune levels for both queens and workers.
- Use data-driven queen selection and handle developmental windows gently to lower epigenetic stress.
| Practice | Benefit | Practical tip |
|---|---|---|
| Forage diversity | Improved immunity, balanced nutrition | Plant mixed bloom patches and avoid monocultures |
| Minimize chemical timing | Lower queen and worker exposure | Apply miticides off-season; use targeted treatments |
| Stress-reduced rearing | Stable offspring performance | Standardize grafting, avoid overheating and crowding |
“Monitor behavioral and reproductive metrics as phenotypic readouts of colony health.”
Collaborate with researchers. Region-specific data on crop landscapes, pesticide profiles, and honey production will refine best practices for U.S. beekeeping and protect bees at the colony and species levels.
Conclusion
A unified picture emerges, where chromatin marks and small RNAs convert social and nutritional cues into lasting gene programs and flexible adult roles.
Gene‑body methylation, histone acetylation, and ncRNA scaffolds act as core mechanisms that tune dna signals and expression to shape growth, reproduction, and behavior. Reversible methylation states then permit rapid task shifts that keep the colony adaptable.
These heritable yet plastic marks connect individual outcomes to population-level change and offer practical levers for U.S. beekeepers: improved nutrition, careful chemical timing, and breeding that limits stress. For a practical view on workforce and task allocation see bee behavior basics.
The role of this framework is testable and actionable. Continued multi‑omic, time‑resolved, cell‑specific work will refine conservation and management strategies for this species.
FAQ
What role does epigenetic regulation play in determining whether a larva becomes a queen or a worker?
Diet-driven chemical signals modify gene activity without changing DNA sequence. These changes—mainly DNA methylation and histone modifications—alter expression patterns that control growth, reproductive organ formation, and lifespan, steering genetically identical larvae toward queen or worker trajectories.
How does royal jelly trigger molecular changes linked to caste outcomes?
Royal jelly contains proteins (MRJPs), fatty acids like 10‑HDA, and other factors that inhibit histone deacetylases and affect methyltransferase activity. These compounds shift chromatin accessibility and methylation states in developing larvae, promoting rapid growth and ovarian development characteristic of queens.
Which DNA methyltransferases are involved and what are their functions?
Two main enzymes mediate methylation dynamics: maintenance methyltransferase DNMT1 preserves existing marks during cell division, while DNMT3 installs new methyl groups. In bees, their coordinated activity shapes tissue‑ and stage‑specific methylation patterns that influence alternative splicing and steady gene output.
Can methylation patterns be tissue-specific or change over time?
Yes. Methylation profiles differ between brain, ovary, and other tissues and shift across larval stages and adult roles. Some marks are stable, while others reverse during behavioral transitions such as nurse-to-forager shifts, reflecting flexible regulation rather than fixed inheritance.
What is the link between methylation and alternative splicing in larvae?
Gene-body methylation frequently occurs across exons and correlates with exon usage. Changes in methylation can modulate splice site choice, producing isoforms with distinct functions that support caste‑specific physiology and neural development.
How do non-coding RNAs contribute to caste-related gene regulation?
Long noncoding RNAs and small RNAs (miRNAs/siRNAs) act as regulators of transcription, mRNA stability, and splicing. Specific lncRNAs expressed in ovaries and brains influence developmental pathways and sensory gene networks, complementing DNA methylation and histone signals.
Are epigenetic marks heritable across generations in bees?
Some methylation sites show conservation across patrilines and can persist through cell lineages, but transgenerational inheritance is limited. Environmental stress and colony context can overwrite marks, so stability varies with genomic location and selective pressures.
Do pesticides or nutrition affect epigenetic programming relevant to beekeeping?
Yes. Chemical exposures and poor nutrition can disrupt methylation and chromatin modifiers, impairing queen production, reducing longevity, and altering immune and metabolic networks. Managing diet quality and limiting harmful chemicals helps preserve healthy epigenetic states.
How reversible are epigenetic changes tied to adult behavioral roles?
Many adult changes are reversible. For example, methylation patterns associated with foraging can revert when individuals return to nursing tasks. This plasticity enables rapid adaptation of brain gene expression to colony needs.
What experimental methods reveal these molecular patterns?
Researchers use whole‑genome bisulfite sequencing for methylomes, ChIP‑seq for histone marks, RNA‑seq for transcriptomes and isoforms, and behavioral reversion models to link molecular changes with function. Integrating these omics approaches clarifies causal relationships.
Which genes and pathways are repeatedly linked to caste differences?
Pathways for insulin signaling, juvenile hormone response, nutrient sensing, and vitellogenesis recur in studies. Gene families like MRJPs and the yellow family, plus regulators of ovary development and metabolism, show consistent caste‑biased expression.
Are there controversies or unresolved questions in this field?
Yes. Researchers debate the extent to which methylation directly controls expression versus reflecting other regulatory states. The specific molecular targets of royal jelly components and the mechanisms linking methylation to behavior remain active research areas.
How does colony-level social environment interact with molecular mechanisms?
Social cues—pheromones such as queen mandibular pheromone, brood signals, and worker feeding behavior—modify larval nutrition and hormonal milieu. These upstream inputs integrate with methylation and chromatin changes to produce coordinated group‑level outcomes.
What practical advice should be applied from epigenomic insights for US beekeepers?
Prioritize balanced nutrition for colonies, reduce exposure to known epigenome‑disrupting agrochemicals, and manage rearing conditions to support queen health. Monitoring brood quality and adopting sustainable feeding practices help maintain favorable gene‑regulatory states.
How do histone modifications complement DNA methylation in regulating gene activity?
Histone acetylation and methylation change chromatin openness and recruit transcriptional machinery. Inhibition of histone deacetylases by royal jelly components increases transcriptional permissiveness, working together with methylation to set developmental trajectories.




